How I Made Python Code 99% Faster with Cython
Overview
Python is a really simple and flexible language when it comes to programming. There’s support for OOP paradigms, functional programming and much more and because of this, it is ubiquitous across various fields such as machine learning, backend development, computer vision, etc. But one qualm developers have when using Python in production-level deployments is that Python can get really slow, really fast.
There are multiple ways to speed up python code. One straight-forward way is to just use a more efficient algorithm. For example, take the famous problem of Two Sum. The naive approach has a time complexityO(N^2) whereas an optimized one takes O(N). But what if there is no better algorithm and we need some other way to speed up our code ?
This specific problem can often be solved by peeping into the programming language itself and finding heuristics to optimize certain paths taken by our program. This is exactly what a JIT compiler such as PyPy does. We won’t delve too deep into this in this post but instead take a different approach to solving our problem.
We all know that low-level languages such as C/C++/Rust/etc are way more performant that high-level languages such as Python … so what if we implement our “time-consuming” functions in any low-level language and .. just call that ? That’s the main idea behind Cython.
Cython is a superset of Python that adds optional, C-like syntax and type annotations, allowing you to write code that looks like Python but can be compiled into fast C extensions. It uses.pyx extension for source files and .pyd files for header files. That’s … pretty cool right ? All we need to do is write our Python implementation in a Python-like language and be done with it ? Right ? …. Well, yes but there is one additional thing that is required. Since C is now involved, there will be a compilation step somewhere down the line. So, we need to set few things up.
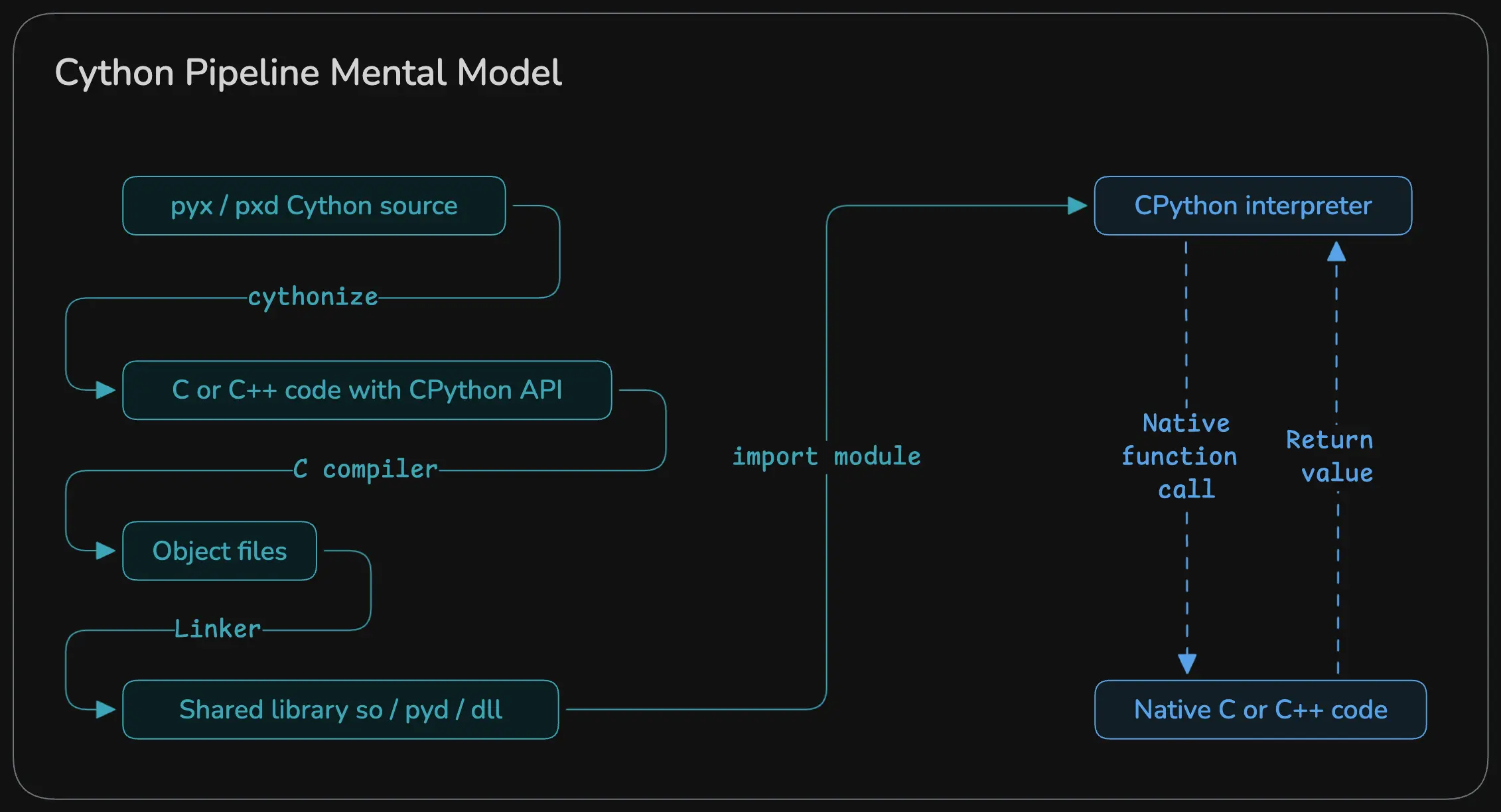
But before that, here’s a general overview of how Cython works under the hood :
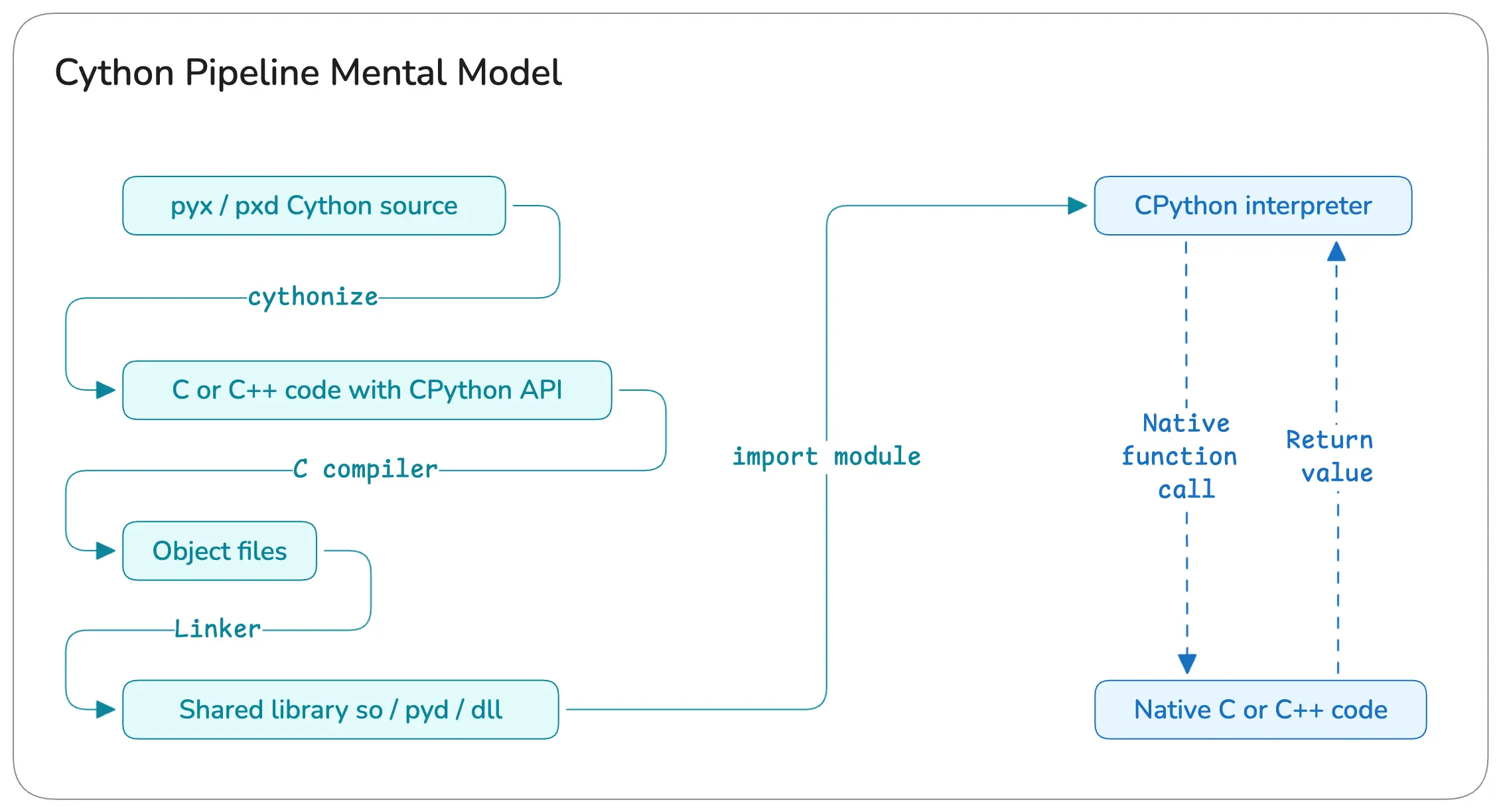
What Are We Optimizing In This Post ?
We will create a simple Matrix class with operations like element wise add / subtract and matmul in Python, port that to Cythonas a pip installable library, benchmark them and see how well each of those perform.
Source code is available on GitHub but it’s heavily under-documented. So, please refer back to this guide till I update that README files there
But First, The Dreaded Setup
There are many tools available but for the sake of simplicity, I will be using uv primarily because it’s fast and snappy. But do note that the uv’s build backend currently supports ONLY python code (see this) so to get around that we will be using setuptools as our build backend
If you want to develop using C / C++ / Cython / Fortran / etc and you prefer a CMake build setup, scikit-build would be better suited for your use case. I won’t be going over that for the sake of simplicity but please check that out if you need something more “concrete”
To scaffold our project (called peak_utils 😎), we will use the following command :
1uv init --build-backend=setuptools peak_utils
This will create a pyproject.toml file something like this : (I added the description so ignore that)
1[project]
2name = "peak-utils"
3version = "0.1.0"
4description = "A lightweight experimental matrix library built with Cython to explore how C/Python interops, build systems, and performance tuning"
5readme = "README.md"
6authors = [
7 { name = "Vishvam10", email = "s.vishvam1025@gmail.com" }
8]
9requires-python = ">=3.14"
10dependencies = []
11
12[project.scripts]
13peak-utils = "peak_utils:main"
14
15[build-system]
16requires = ["setuptools>=61", "wheel", "Cython"]
17build-backend = "setuptools.build_meta"
Now, we need to somehow call the cython compiler to convert our .pyx / pxd code to .c code. The simplest way to do that is to include that in a special script called setup.py. This is a build script for Python projects. Think of it like a Makefile / CMakeLists.txt, but for Python packaging.
We have declared Cython as a required library under [build-system] so it’s readily available for us to use without doing uv add Cython. If you are using pip, please create a virtual environment and do pip install Cython setuptools wheels.
1from setuptools import setup, Extension, find_packages
2from Cython.Build import cythonize
3
4# We can add more files here if needed. IIRC there is also a glob support for this
5# but don't quote me on that 😝
6extensions = [
7 Extension(
8 "peak_utils.matrix",
9 ["src/peak_utils/matrix.pyx"],
10 ),
11]
12
13setup(
14 name="peak_utils",
15 version="0.1.0",
16 package_dir={"": "src"},
17 packages=find_packages(where="src"),
18 ext_modules=cythonize(extensions)
19)
Most of the fields here are quite self-explanatory so I’m not going to go over each of them. Just note the package_dir and packages key. We want to get the source files from the src folder. And that’s basically it. For this example, we need only these 2 files to set things up. If you are following this as a guide, then make sure this is the directory structure you have :
1peak_utils
2├── README.md
3├── pyproject.toml
4├── setup.py
5├── src
6│ └── peak_utils
7│ ├── __init__.py
8│ ├── matrix.c
9│ ├── matrix.pxd
10│ └── matrix.pyx
11└── uv.lock
Now, Let’s Code
This is the Python implementation that we will be porting to Cython :
1class PyMatrix:
2 def __init__(self, rows, cols):
3 self.rows = rows
4 self.cols = cols
5 self.data = [0.0] * (rows * cols)
6
7 @classmethod
8 def from_list(cls, values):
9 r = len(values)
10 if r == 0:
11 raise ValueError("Empty list")
12 c = len(values[0])
13
14 m = cls(r, c)
15 for i in range(r):
16 if len(values[i]) != c:
17 raise ValueError("Jagged list")
18 for j in range(c):
19 m.data[i * c + j] = float(values[i][j])
20 return m
21
22 def add(self, other):
23 if self.rows != other.rows or self.cols != other.cols:
24 raise ValueError("Shape mismatch")
25
26 out = PyMatrix(self.rows, self.cols)
27 for i in range(self.rows * self.cols):
28 out.data[i] = self.data[i] + other.data[i]
29 return out
30
31 def sub(self, other):
32 if self.rows != other.rows or self.cols != other.cols:
33 raise ValueError("Shape mismatch")
34
35 out = PyMatrix(self.rows, self.cols)
36 for i in range(self.rows * self.cols):
37 out.data[i] = self.data[i] - other.data[i]
38 return out
39
40 def matmul(self, other):
41 if self.cols != other.rows:
42 raise ValueError("Shape mismatch")
43
44 out = PyMatrix(self.rows, other.cols)
45
46 for i in range(self.rows):
47 for j in range(other.cols):
48 s = 0.0
49 for k in range(self.cols):
50 s += (
51 self.data[i * self.cols + k] *
52 other.data[k * other.cols + j]
53 )
54 out.data[i * other.cols + j] = s
55 return out
Nothing too fancy. Just a simple, naive implementation of a matrix. Now, you’ll be surprised how easy it is to port this to Cython. First, we’ll start with header files. This is where we’ll have the definitions :
Header Files
Create a new file called matrix.pxd :
1cdef class Matrix :
2 cdef Py_ssize_t rows
3 cdef Py_ssize_t cols
4 cdef double* data
5
6 cpdef Matrix add(self, Matrix other)
7 cpdef Matrix sub(self, Matrix other)
8 cpdef Matrix matmul(self, Matrix other)
cdef basically means “this method / class is accessible by C code”, def, well that’s our regular python function and cpdef means “this method is accessible by C and Python code. Now, you might wonder why not use int or something for the number of rows and cols. The reason for usingPy_ssize_t is that, this is will always resolve to the platform-specific size of an integer
Cython Compiler Directives
Create a new file called matrix.pyx and define a few crucial information for the cython compiler : (the # symbol is required and is used as a compiler directive)
1# cython: language_level=3
2# cython: boundscheck=False, wraparound=False, nonecheck=False
3
4from libc.stdlib cimport malloc, free
5import cython
language_level=3: sincecythonpredatesPython 3, we specifically ask thecythoncompiler to usePython 3semantics such as :- / is true division
- print() is a function
- Strings are unicode by default, etc
boundscheck=False:- Disables array bounds checking for :
C arraysandtyped memoryviews(more on this later) - Why it matters : It removes branching, enables vectorization and much faster inner loops
- Disables array bounds checking for :
1# Example without this directive :
2x = A[i, j]
3
4# Cython normally generates :
5
6if (i < 0 || i >= A.shape[0]) error;
7if (j < 0 || j >= A.shape[1]) error;
8
9# With this directive, it becomes :
10x = A_data[i * stride0 + j * stride1];
wraparound=False: Disables Python-style negative indexing (soarr[-1]is no longer valid)- Why it matters : fewer checks, faster indexing
nonecheck=False: Disables checks likeif A is None : ...- Why it matters : same reason as above, fewer checks (as it doesn’t need to evaluate this :
if (A == Py_None) error;so has a faster attribute access and also less branching
- Why it matters : same reason as above, fewer checks (as it doesn’t need to evaluate this :
All these performance boosts comes with a cost that programmers have to keep in mind :
- By chance if you get an out-of-bounds error, that becomes a Segfault, not Python exception.
- By chance if you pass something as
Nonewhich is NOT supposed to be aNone, the program will crash
from libc.stdlib cimport malloc, free:- We use
cimportto import C libraries. This basically imports the C functionsmallocandfreefrom the C standard library so that we can call them directly from Cython-generated C code
- We use
import cythonis a compile-time helper module that gives us access to things like : (these are not available in the Python runtime)
1cython.view.array(...)
2cython.boundscheck(False)
3cython.cdivision(True)
4cython.inline
5cython.final
On To The Implementation Part
First and foremost, we need to manage memory when Matrix is created and destroyed. Cython provides 2 handy functions : __cinit__ and __dealloc__ . On the same matrix.pyx file, let’s create the Matrix class :
1# cython: language_level=3
2# cython: boundscheck=False, wraparound=False, nonecheck=False
3
4from libc.stdlib cimport malloc, free
5
6cdef class Matrix :
7
8 # For initializing and managing memory
9
10 def __cinit__(self, Py_ssize_t rows, Py_ssize_t cols) :
11 self.rows = rows
12 self.cols = cols
13 self.data = <double*> malloc(rows * cols * sizeof(double))
14
15 if self.data == NULL :
16 raise MemoryError("Matrix must have some data")
17
18 def __dealloc__(self) :
19 if self.data != NULL :
20 free(self.data)
21
22 # For printing and debugging
23
24 def __str__(self):
25 cdef Py_ssize_t i, j
26 lines = []
27 for i in range(self.rows):
28 row = []
29 for j in range(self.cols):
30 row.append(str(self.data[i * self.cols + j]))
31 lines.append("[ " + ", ".join(row) + " ]")
32 return "\n".join(lines)
33
34 def __repr__(self):
35 return f"Matrix({self.rows}, {self.cols})\n{self.__str__()}"
36
37 @classmethod
38 def from_list(cls, list values):
39 cdef Py_ssize_t r = len(values)
40 if r == 0:
41 raise ValueError("Empty list")
42
43 cdef Py_ssize_t c = len(values[0])
44 cdef Matrix m = cls(r, c)
45 cdef Py_ssize_t i, j
46
47 for i in range(r):
48 if len(values[i]) != c:
49 raise ValueError("Jagged list")
50 for j in range(c):
51 m.data[i * c + j] = values[i][j]
52 return m
<Type*>is the syntax used byCythonto do type casting, here we mention that the memory received frommallocwill store a bunch ofdouble(s). More on that here@classmethodis the same one that we use inPython. Please refer to this link to know more about that
Now, let’s implement the operations :
1# cython: language_level=3
2# cython: boundscheck=False, wraparound=False, nonecheck=False
3
4from libc.stdlib cimport malloc, free
5
6cdef class Matrix :
7
8 ...
9
10 cpdef Matrix add(self, Matrix other) :
11
12 if self.rows != other.rows or self.cols != other.cols:
13 raise ValueError("Shape mismatch")
14
15 cdef Matrix out = Matrix(self.rows, self.cols)
16 cdef Py_ssize_t i, n = self.rows * self.cols
17
18 for i in range(n) :
19 out.data[i] = self.data[i] + other.data[i]
20
21 return out
22
23
24 cpdef Matrix sub(self, Matrix other) :
25
26 if self.rows != other.rows or self.cols != other.cols:
27 raise ValueError("Shape mismatch")
28
29 cdef Matrix out = Matrix(self.rows, self.cols)
30 cdef Py_ssize_t i, n = self.rows * self.cols
31
32 for i in range(n) :
33 out.data[i] = self.data[i] - other.data[i]
34
35 return out
36
37
38 cpdef Matrix matmul(self, Matrix other) :
39
40 if self.cols != other.rows:
41 raise ValueError("Shape mismatch")
42
43 cdef Matrix out = Matrix(self.rows, other.cols)
44 cdef Py_ssize_t i, j, k
45
46 for i in range(self.rows):
47 for j in range(other.cols):
48 out.data[i * other.cols + j] = 0.0
49 for k in range(self.cols):
50 out.data[i * other.cols + j] += (
51 self.data[i * self.cols + k] *
52 other.data[k * other.cols + j]
53 )
54
55 return out
Note that we didn’t make any change to our core (and naive) algorithm, we just adapted Cython syntax. We could annotate every method’s return type and other things to squeeze in more performance but for the time being this is fine.
Building The Library
Oh boy, this is where it took me some time to figure things out. If you get stuck while building using uv, please refer to this GitHub thread. For this specific guide, this is the folder structure that we are following :
1tree -I '.git|.venv|__pycache__'
2.
3├── README.md
4├── benchmark
5│ ├── README.md
6│ ├── bench.py
7│ ├── benchmark_results.csv
8│ ├── plot.py
9│ ├── pymatrix.py
10│ ├── pyproject.toml
11│ └── uv.lock
12└── peak_utils
13 ├── README.md
14 ├── pyproject.toml
15 ├── setup.py
16 ├── src
17 │ ├── peak_utils
18 │ │ ├── __init__.py
19 │ │ ├── matrix.c
20 │ │ ├── matrix.pxd
21 │ │ └── matrix.pyx
22 │ └── peak_utils.egg-info
23 │ ├── PKG-INFO
24 │ ├── SOURCES.txt
25 │ ├── dependency_links.txt
26 │ ├── entry_points.txt
27 │ └── top_level.txt
28 └── uv.lock
To build the library, go into peak_utils (top one) and run uv build :
1 cd peak_utils
2 uv build --wheel
wheel is a standard binary distribution format for Python packages. There are other things such as sdist (which is basically metadata + tar of source code), egg, etc but we won’t go too deep into that. Given that this post is long enough, I’ll have a separate write-up for this 😅
Running this command should ideally give an output such as this :
1.. a bunch of 'writing', 'building' stuff
2Successfully built dist/peak_utils-0.1.0-cp314-cp314-[your arch].whl
and you should have a build and a dist folder
Benchmarking
To use our newly created package, run the following command on benchmark folder :
1 cd benchmark
2 uv sync
3
4 uv add ../peak_utils/dist/peak_utils-0.1.0[your arch].whl
The code for benchmarking and plotting are mostly boilerplate so I’m just gonna link that here :
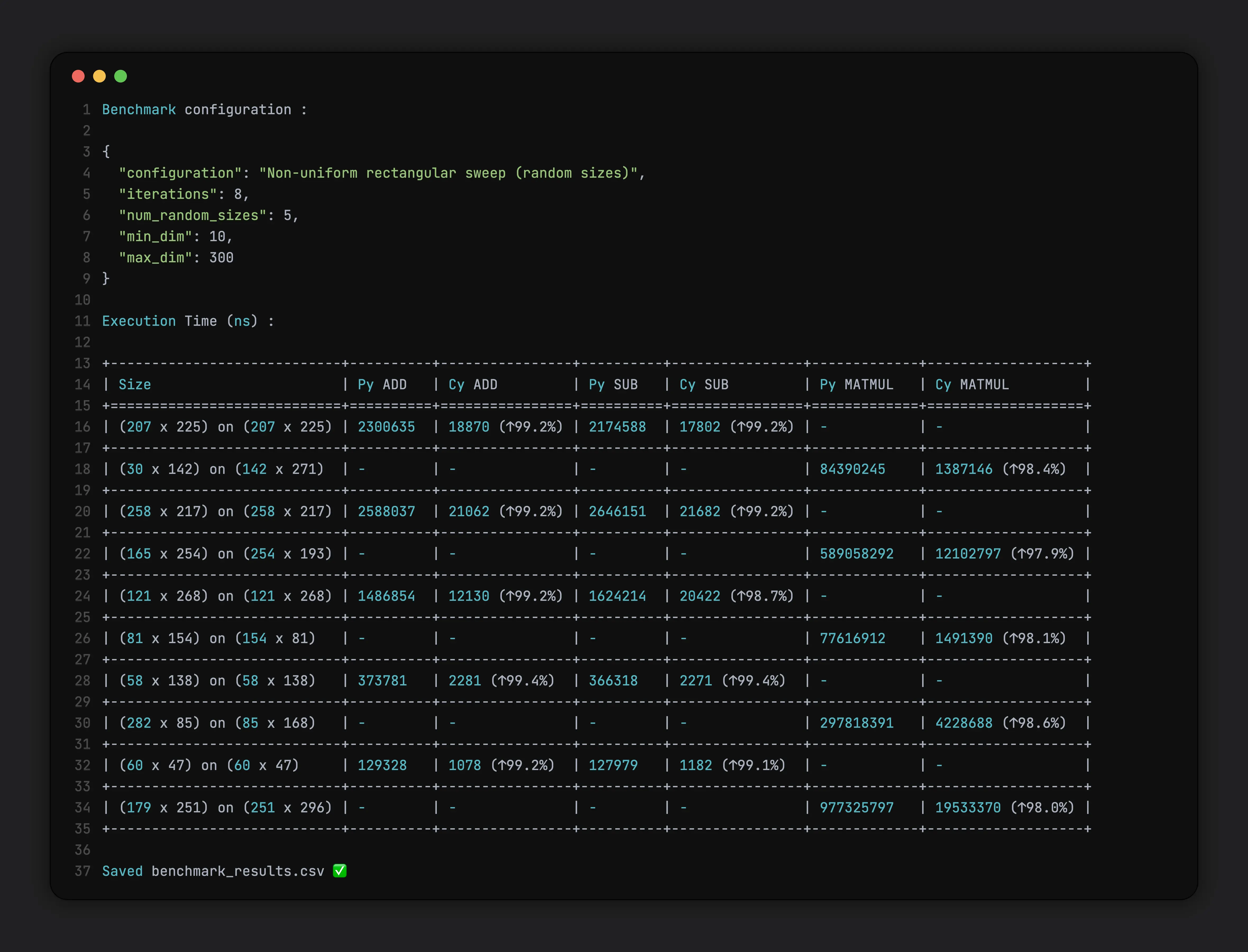
We will use these params :
1{
2 "configuration": "Non-uniform rectangular sweep (random sizes)",
3 "iterations": 8,
4 "num_random_sizes": 5,
5 "min_dim": 10,
6 "max_dim": 300
7}
where
configurationis either a uniform (N x N) square matrices OR non-uniform (N x M) rectangular matricesiterationsis the number of runs we do (to get mean, variance, etc)num_random_sizesis the number of matrices createdmin_dim and max_dimare the min and max dimensions of matrices (randomly chosen)
Results And Pretty Plots
The benchmarks were conducted on an Apple Macbook Air with an M3 Chip, 16GB RAM and 512GB storage
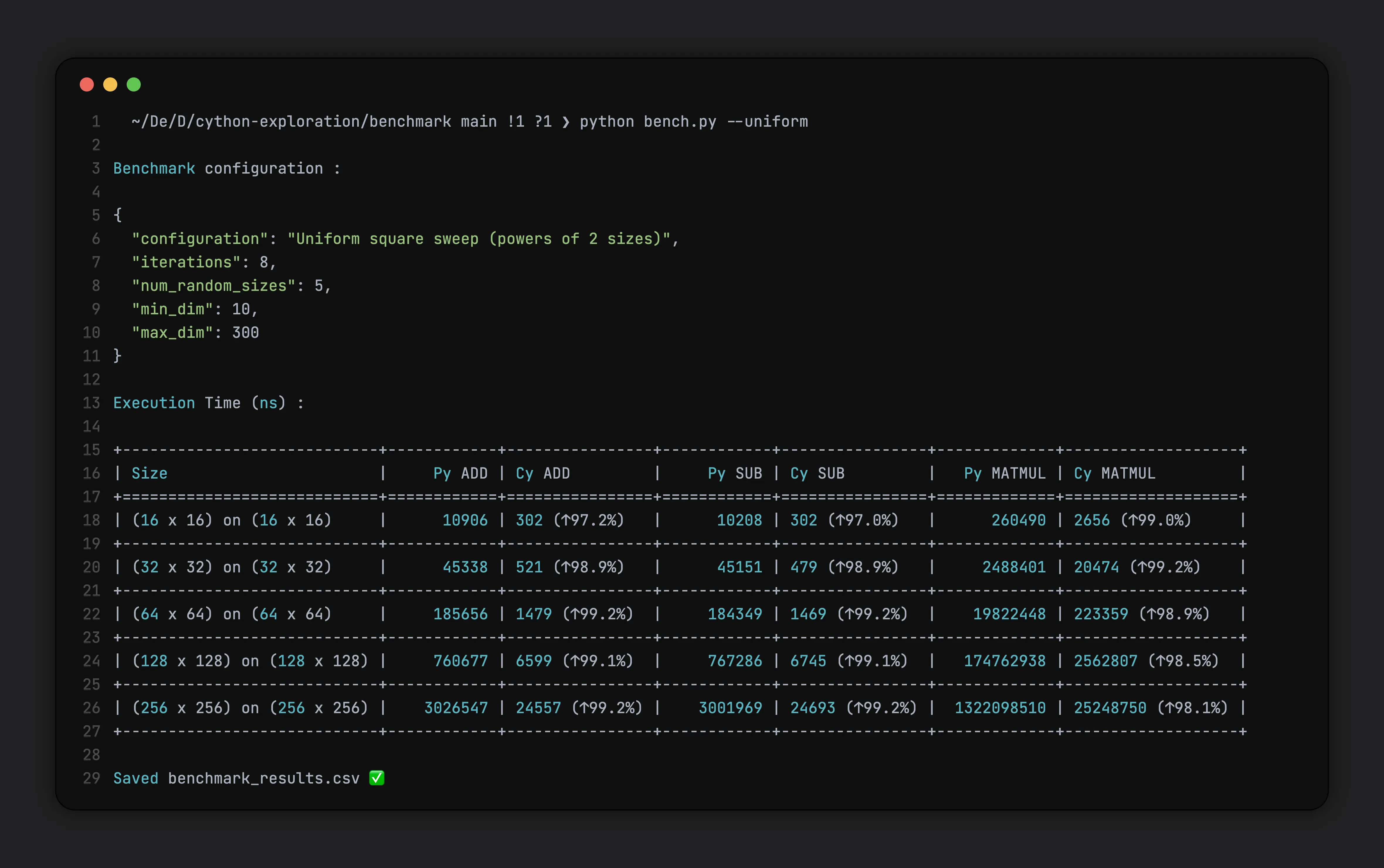
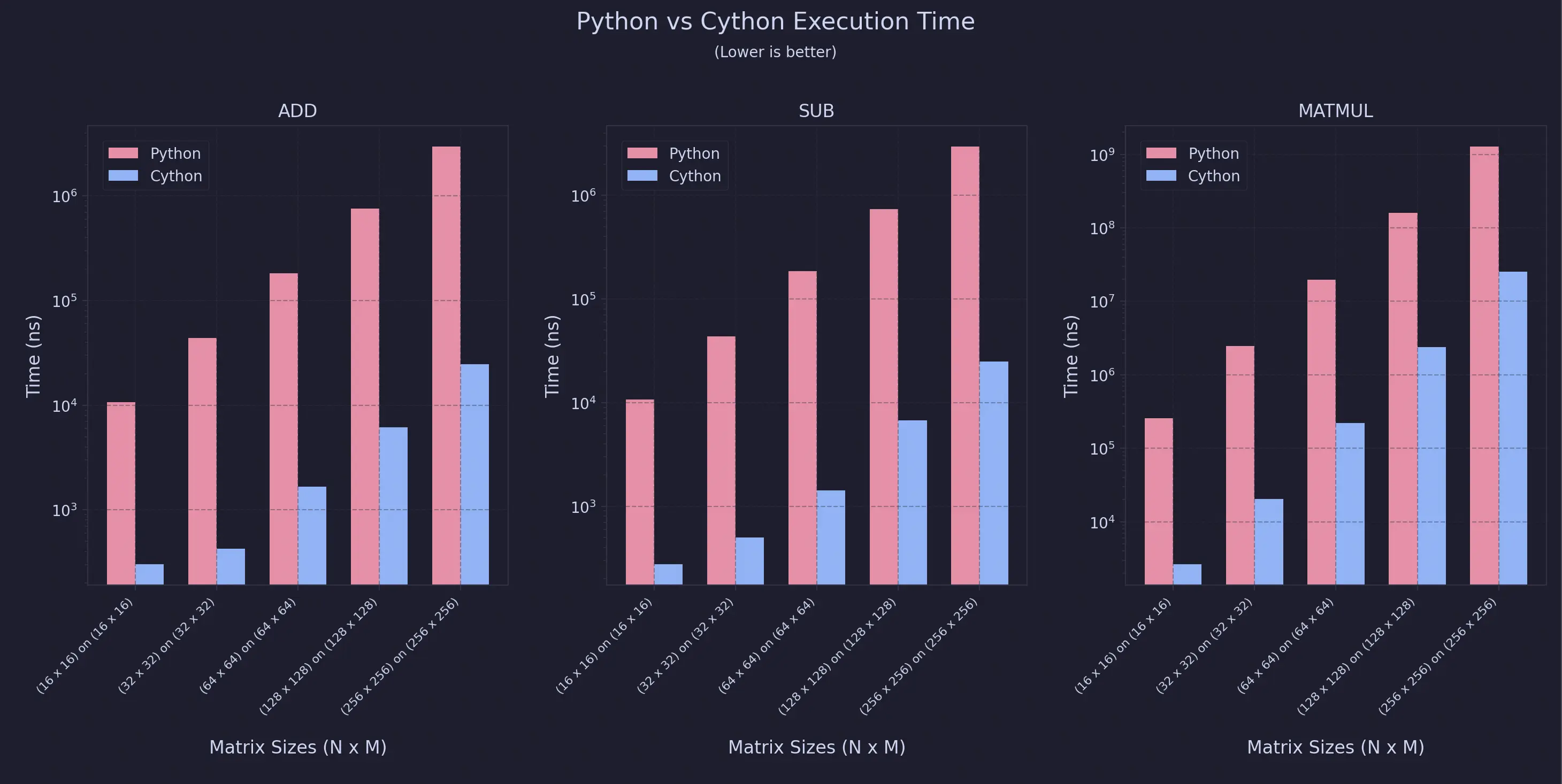
For Uniform Sweeps
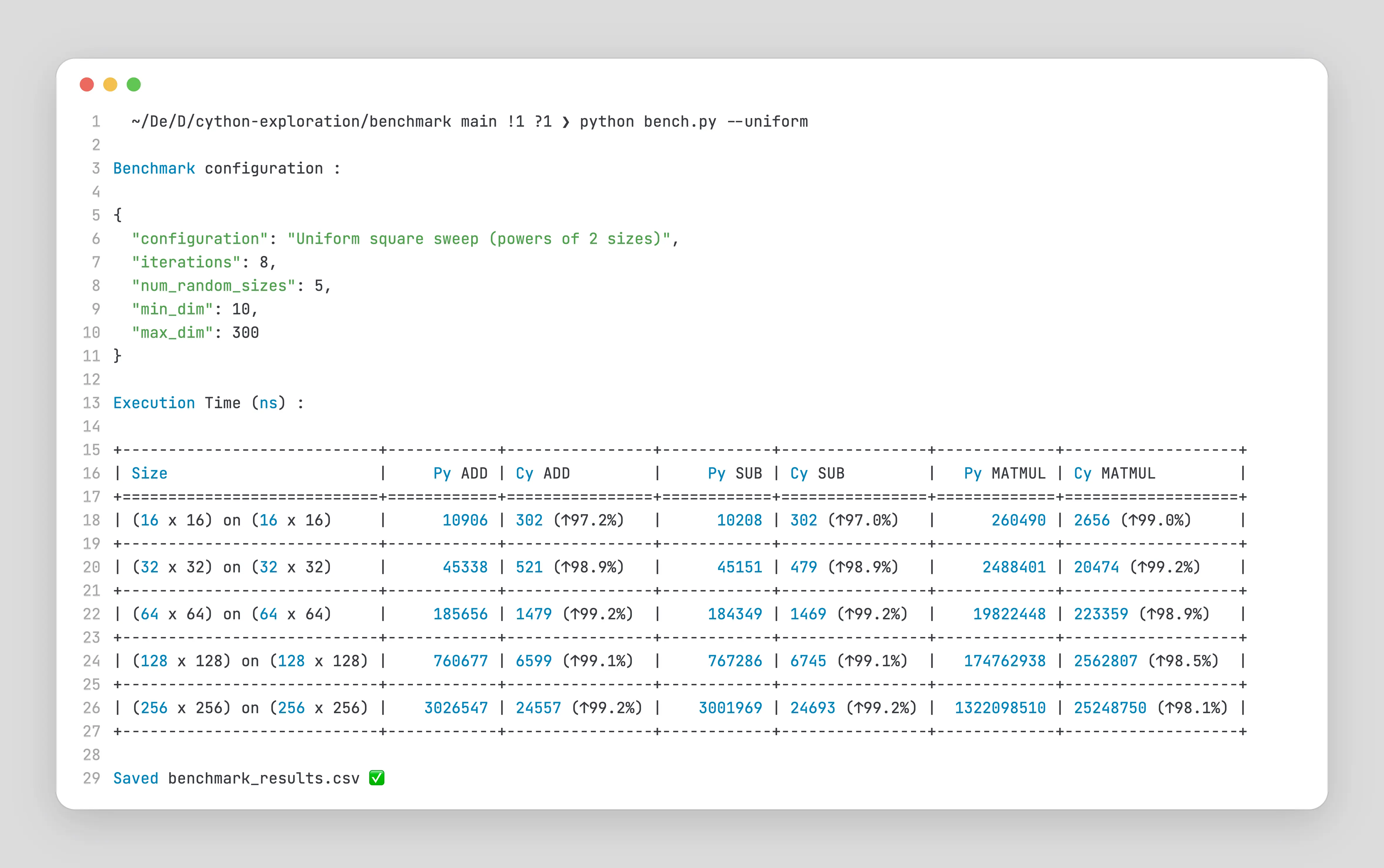
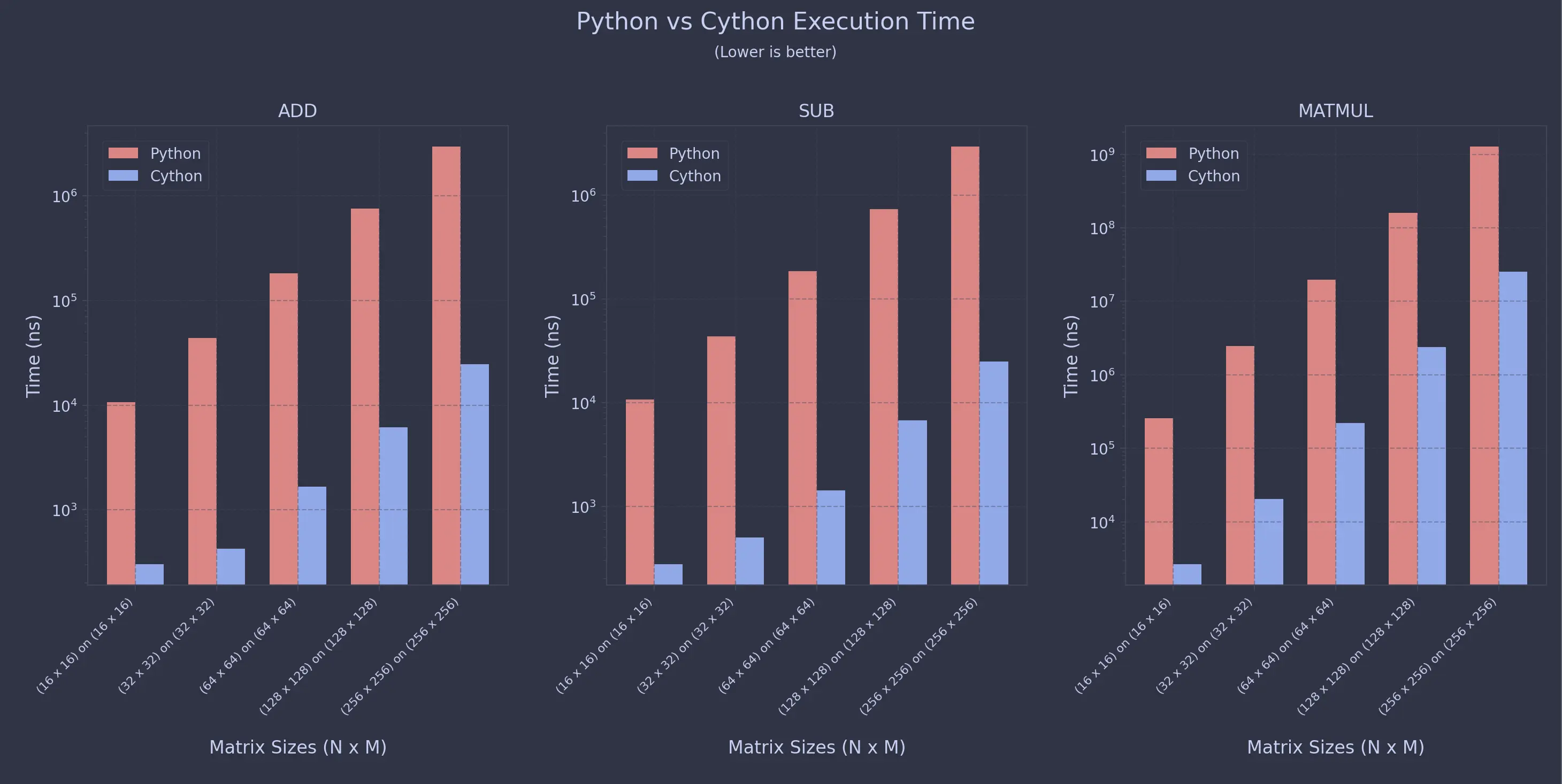
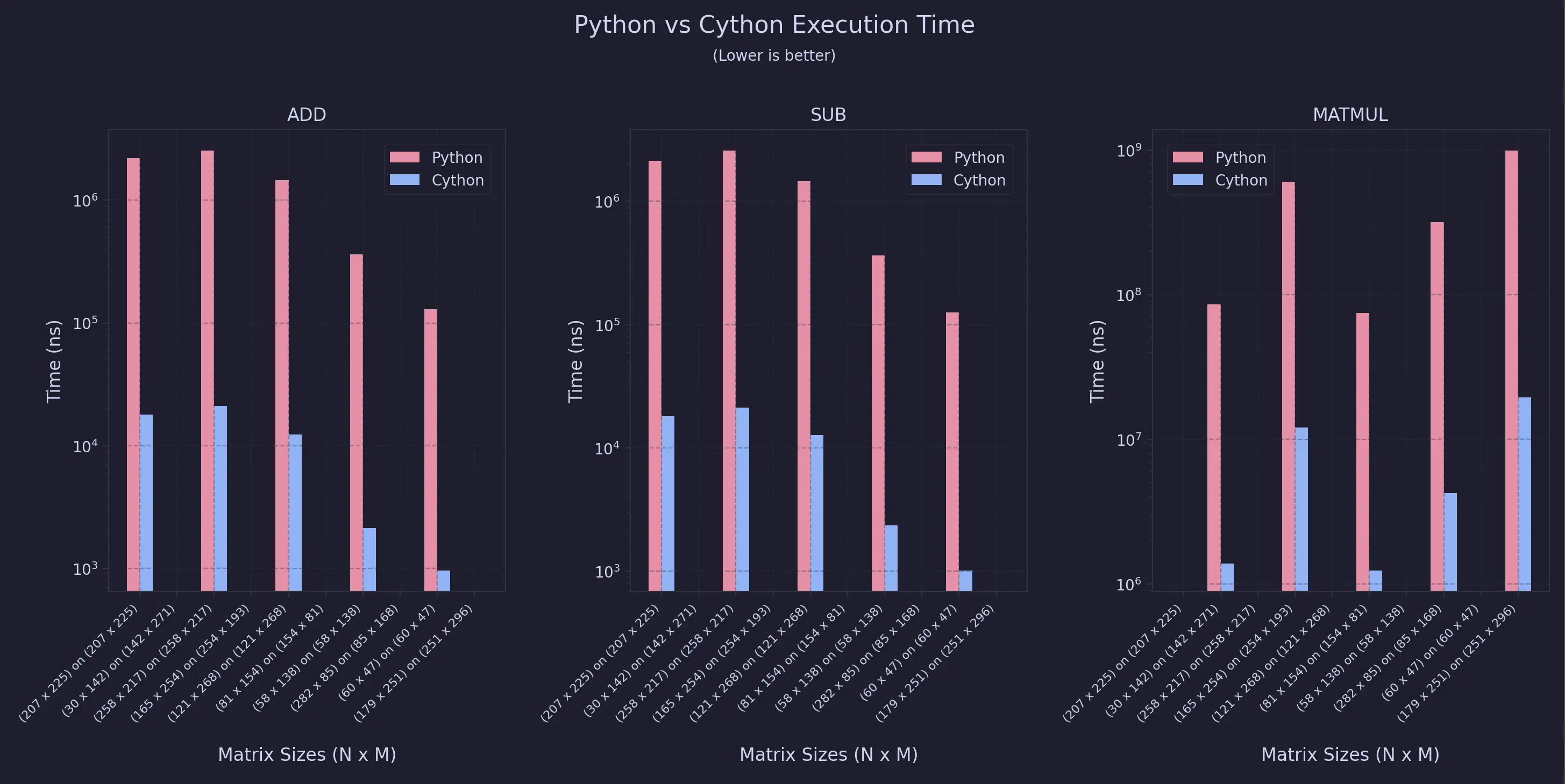
For Non-Uniform Sweeps
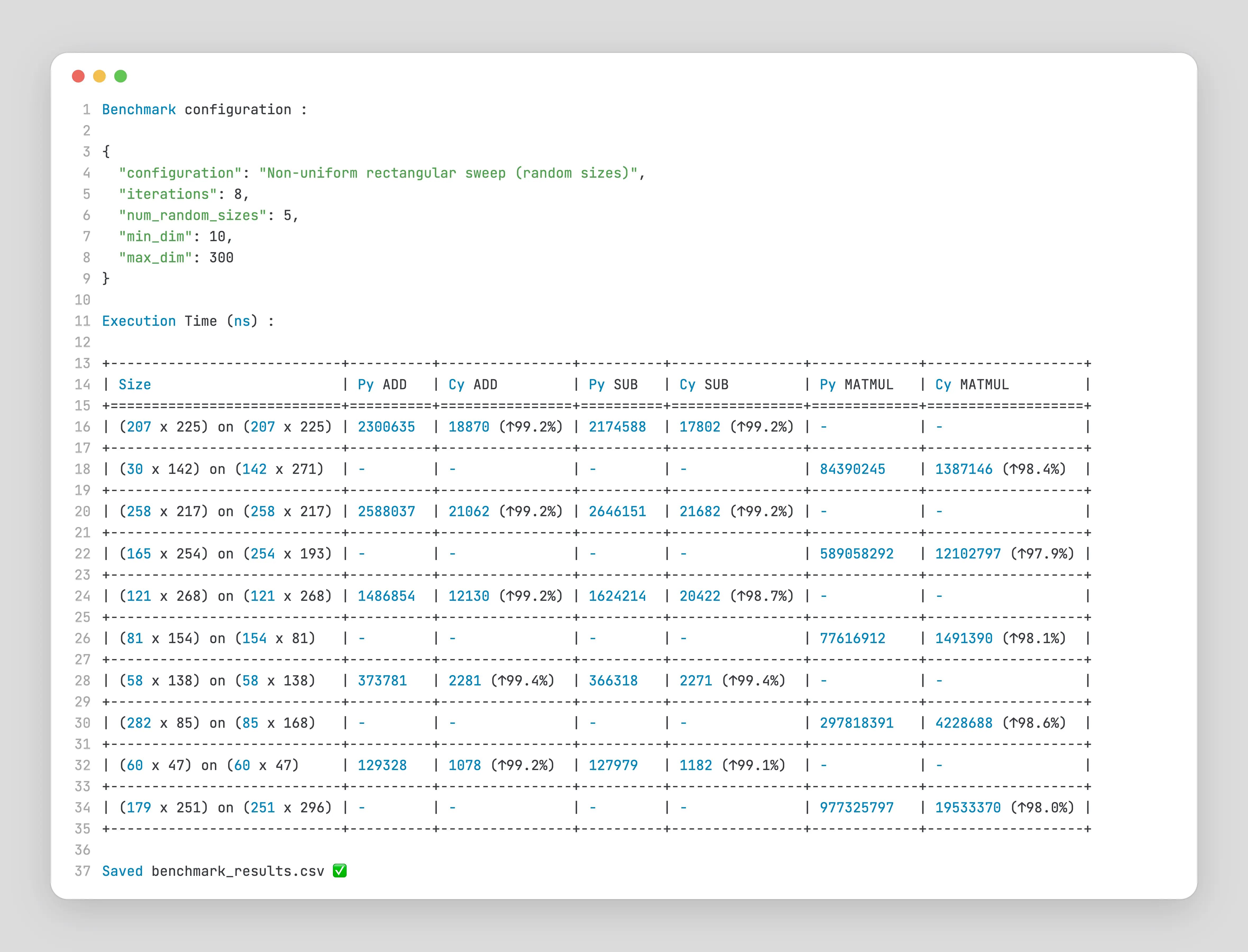
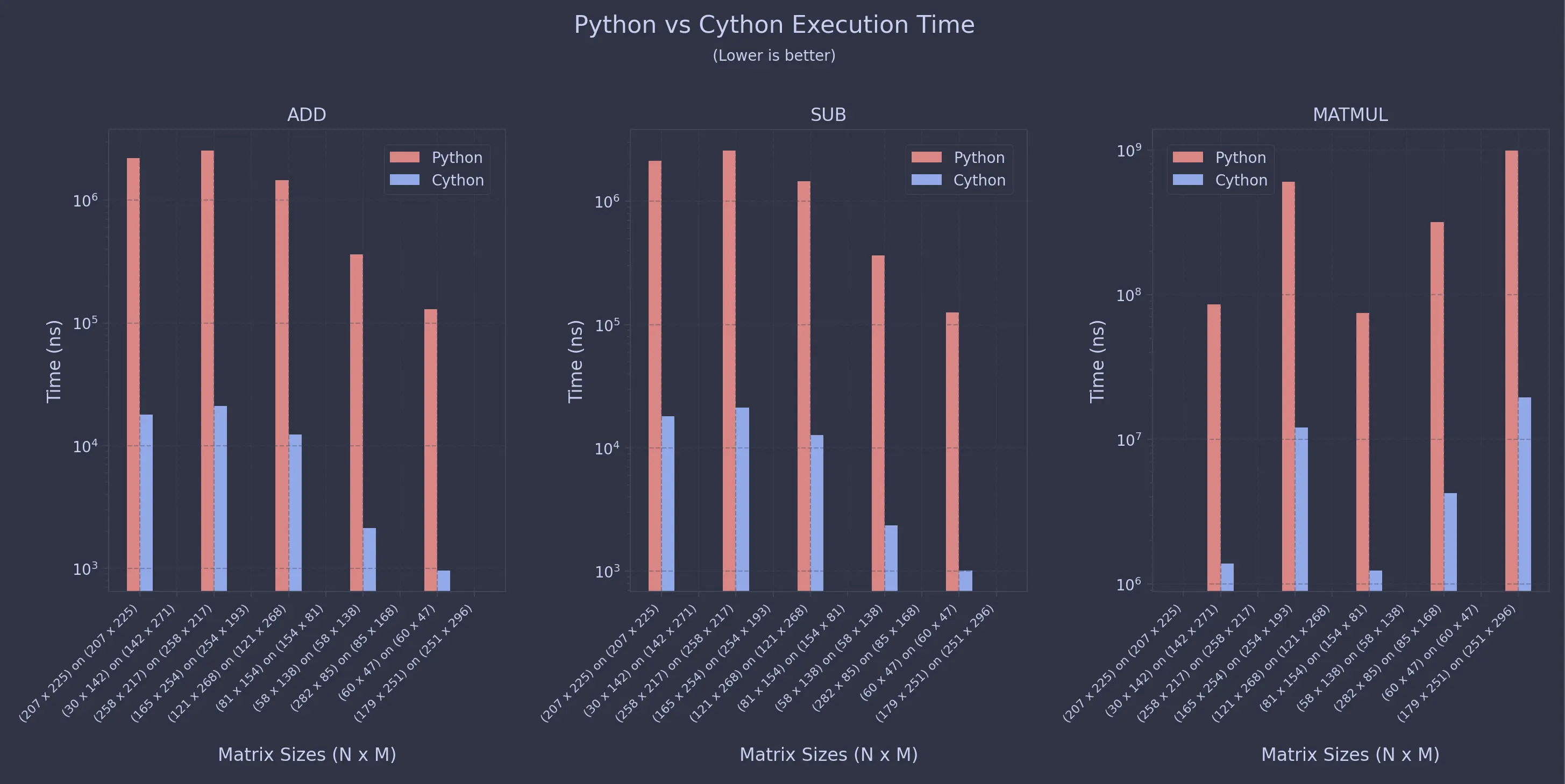
As you can see, the speedup is almost always 99% for this specific configuration.
Conclusion
So what’s the real takeaway here ? It’s not just that Cython is fast, it’s why it’s fast. Once you move those tight inner loops out of Python’s runtime and into compiled C, a lot of the usual overhead just disappears. You still get to write code that feels like Python, but it runs much closer to native speed.
Of course, this isn’t a magic button. There’s extra setup, a build step, and now you’re also responsible for things like memory management and not shooting yourself in the foot with segfaults. In a lot of cases, using better algorithms, NumPy, or even PyPy will get you most of the way there. But when you’ve really hit the ceiling and Python itself is the bottleneck, Cython is a pretty sweet middle ground
And if you’re willing to get your hands dirty, you can push things even further: static typing, raw memory buffers, bounds-check removal, OpenMP parallelism - some crazy optimizations can take your speedups well beyond what typical Python code ever sees.
Comments